Before you read this, I suggest you read post 21.2.
Most people who are interested in science know that the DNA molecule is a double helix. But what does this mean? How do we know? And why is it important?
By the early 1950s, there was evidence that genes were made of DNA and that the sequences of its bases must code this genetic information (see post 21.11). Following Pauling’s discovery of the α-helix, several people wondered whether the three-dimensional structure of the DNA molecule might be important for its genetic function. The experimental evidence that led to the discovery of the α-helix was provided by x-ray fibre diffraction. The first x-ray fibre diffraction patterns of DNA were obtained by Astbury. However, the experimental results that led to the discovery of the DNA double helix were obtained by two independent research groups at King’s College London: one led by Rosalind Franklin, the other by Maurice Wilkins. Pauling had interpreted x-ray diffraction results by molecular model building (see post 21.7).
Francis Crick and James Watson, at the University of Cambridge, decided to use the same approach to determining the three-dimension structure of the DNA molecule. Following Pauling, they reasoned that a repetitive molecule like DNA was likely to coil into a helix. But what would this helix look like?
X-ray diffraction showed that the axial rise-per-residue of the helix would be 0.34 nm and its pitch would be 3.4 nm; so the molecule would be an integral 10-fold helix (because 3.4/0.34 = 10, all this is explained in post 21.7). Crick and Watson also incorporated some further ideas into their modelling.
- The number of adenine (A) bases must be equal to the number of thymine (T) bases and the number of guanine (G) bases must be equal to the number of cytosine (C) bases (see post 21.2).
- The phosphate groups were hydrophilic and so likely to be on the outside of the helix.
- The bases were hydrophobic and so likely to be on the inside of the helix, protecting them from surrounding water molecules.
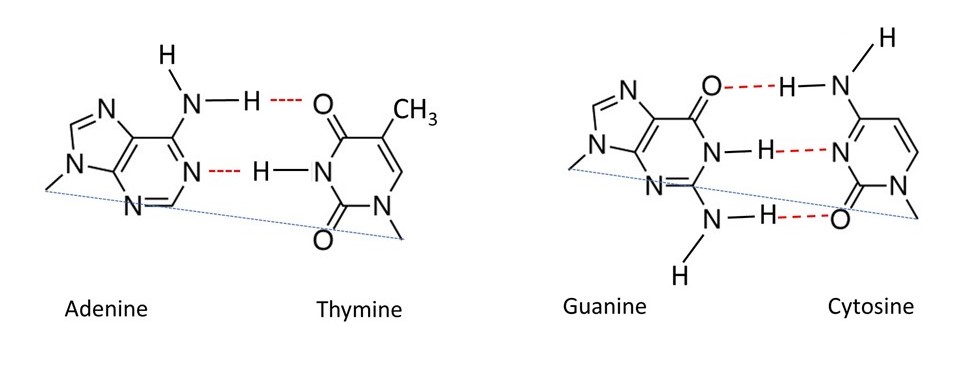
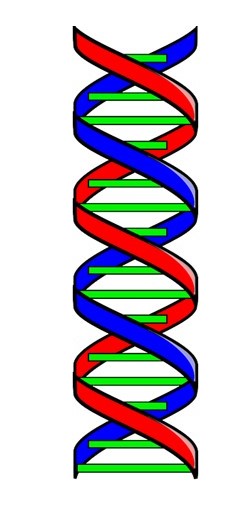
They first showed that A-T and G-C pairs of bases could be held together by hydrogen bonds (shown by red dashed lines in the pictures above) that had the same overall dimensions as each other. In the pictures the ends of the bonds that join the nitrogen atoms (in each base pair) to sugar residues are joined by a blue dotted line. In both A-T and G-C pairs, these lines are the same length and make the same angles with the bonds. These base pairs could be incorporated into a structure consisting of two coaxial sugar phosphate chains, pointing in opposite directions, represented by the picture below. Here the base pairs are shown in green. Since A-T and G-C pairs have the same overall size and shape, either could occur in any position within this anti-parallel double helix. One sugar-phosphate chain is shown in red, the other in blue: one points “up” the other points “down”. Apart from their directions, the two sugar-phosphate chains can be identical.
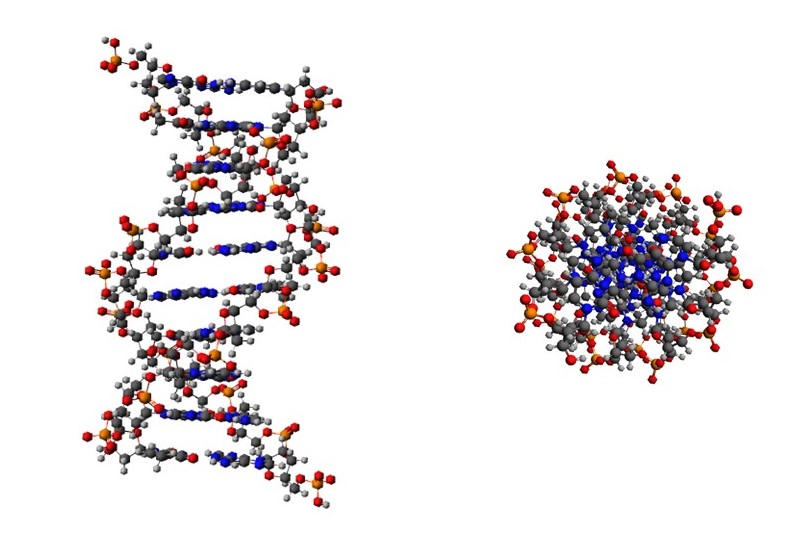
When Crick and Watson started to build models of the sugar phosphate chains, they found it was possible to build right-handed helices, in which non-bonded atoms did not approach more closely than would be possible. And these helices had the pitch and axial rise-per-residue determined by x-ray diffraction. The positions of the atoms, in a length of this double helix are shown in the pictures below. On the left-hand side, we are looking perpendicular to the helix axis. On the right-hand side, we are looking down the helix axis and can see the phosphate groups on the outside and the bases on the inside.
Most accounts of the discovery of the double helix stop here. But further experimental tests were required. These were provided by Wilkins and his colleagues. They improved the original model built by Crick and Watson to show that it could give a good explanation of the x-ray intensity distribution of their diffraction patterns, including computer modelling, as explained in appendix 3 of post 21.7.
So, now we know what it means to say that DNA is a double helix and the evidence for this belief.
But why is the double helix important? It appears that the sequence of bases provides genetic information (post 21.11). Suppose that the sequence of bases on one chain is AGCTTCGA, then the sequence on the other chain must be TCGAAGCT because A always pairs with T and G always pairs with C. The sequence on one chain could code the genetic information and the sequence on the other is complementary to it – and so could be used as a template to form a new coding sequence. The double helix can accommodate any sequence of bases, with its complementary sequence. I will give more information on these ideas in a later post.
In summary, it is the sequence of bases that is important for the genetic function of DNA molecule. The double helix is a structure which can accommodate any genetic information together with the information that allows a copy of this information to be made.
Related posts
21.11 Genes are made of DNA
21.10 Chromosomes 2
21.9 Chromosomes 1
21.8 Genes
21.7 The α-helix
21.6 The helix
21.2 Nucleic acids
Follow-up posts