Before you read this, I suggest you read post 21.23.
In post 21.23 we met the idea that the sequence of bases in mRNA determined the sequence of amino acids in a protein. The question then is – what sequence of bases will code a given amino acid sequence? We shall see that the experiments that determined this genetic code provide further evidence for the ideas in post 21.23.

How many bases are required to code for a single amino acid? There are about 20 different amino acids in proteins but only 4 bases (A, U, G and C) in mRNA. Therefore, more than one base is required to code for an amino acid. Let’s suppose that the genetic code is a two-letter code; the first letter can be chosen in 4 ways and the second can also be chosen in 4 ways, making a total of 4 × 4 = 16 possible combinations, as shown in the table above. This is not enough to code for 20 different amino acids. But, if the genetic code is a three-letter code, each letter can be chosen in 4 different ways, giving 4 × 4 × 4 = 64 different combinations, also shown in the table above. A three-letter code could specify all the amino acids found in proteins.

The next question is whether the code is non-overlapping or overlapping. In a non-overlapping three-letter code, the first three letters code one thing and the next three letters code the next. As an example, think of three-letter words in the English language. We read these letters as a non-overlapping code so that the letters TWONET give the words “two” and “net”. In an overlapping code, the first three letters give the first word (“two”) but the second word (“won”) is given by the second, third and fourth letters. The difference between the two types of code is shown in the picture above.
The problem with an overlapping code is that it restricts choice because, apart from the first word, the choice of word is limited by the words that precede it. You will see the difficulty if you try to find more three-letter English words that can be formed following TWONET in our example of an overlapping code. In the context of the genetic code, this would limit the possible number of proteins that could be made. Since our bodies contain between 80 000 and 400 000 different proteins this could be a serious problem.
So, it seems reasonable to suppose that the genetic code would be a non-overlapping three letter code.
Experimental evidence for this supposition enabled the genetic code to be cracked by Marshall W Nirenberg and his collaborators at the National Institutes of Health in the USA. These experiments were only possible because Har Gobind Khorana, at the University of Wisconsin, had started to make synthetic mRNA with different sequences. The experiments involved cell-free synthesis of proteins in a system containing amino acids, ribosomes, tRNAs, aminoacyl-tRNA synthetases (the enzymes that are catalysts for the attachment of an amino acid to an appropriate tRNA) and ATP (as a source of energy for the chemical reactions involved).
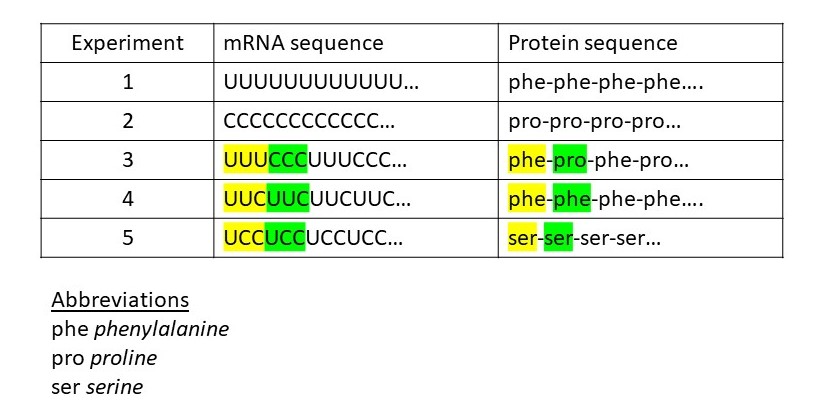
Let’s look at some examples of these experiments (see table above). In experiment 1, we see that mRNA in which all the bases are U codes for poly(phenylalanine). In experiment 2, when all the bases are C, the mRNA codes for poly(proline). Experiment 3 suggests that the code is a non-overlapping three-letter code. Further evidence for this conclusion is provide by experiments 4 and 5 which show that if the code were overlapping, the protein sequence in experiment 3 would be: phe-phe-ser-pro…

Many experiments of this kind were consistent with the non-overlapping three-letter code shown in the table above. In this table, the entries in blue are the punctuation mark STOP which denotes the end of a protein sequence. These cell-free experiments also provided evidence for the ideas about transcription and translation described in post 21.23.
Note that there are several different base sequences that code for the same amino acid – the genetic code is an example of a redundant code.
Some proteins contain amino acids that don’t appear in the table above – examples are hydroxyproline and hydroxylysine. These are formed by chemical modification of proline and lysine residues after translation has occurred; this process is called post-transcriptional modification.
In conclusion, if we know the sequence of bases in mRNA or DNA, we know the sequence of amino acids that they code for, but we can’t tell if they would be modified after transcription.
Related posts
21.8 Genes
21.11 Genes are made of DNA
21.12 The DNA double helix
21.14 Copying genetic information
21.23 Implementing genetic information